pISSN : 3058-423X eISSN: 3058-4302
Open Access, Peer-reviewed
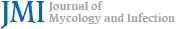
pISSN : 3058-423X eISSN: 3058-4302
Open Access, Peer-reviewed
Tuan Khoi Tran,Waewta Kuwatjanakul,Lumyai Wonglakorn,Kittipan Samerpitak
10.17966/JMI.2026.31.1.25 Epub 2026 March 31
Abstract
Background: ITS barcoding is the "gold standard" method for fungal identification. However, it is associated with high costs, requires specialized expertise, and entails lengthy turnaround times. MALDI-TOF MS represents a more rapid and lower-cost alternative, but it is generally less sensitive and less specific than ITS barcoding.
Objective: This study aimed to investigate and assess MALDI-TOF MS and ITS barcoding for identifying yeast strains isolated from patients and domestic environments.
Methods: A total of 115 yeast strains, including 89 pathogenic and 26 environmental, were identified using MALDI-TOF MS and ITS barcoding with phylogenetic analyses. The identification results from the two methods were compared and assessed.
Results: ITS barcoding identified 115 strains, including ascomycetous yeasts (5 Candida species and 9 non-Candida species) and basidiomycetous yeasts (5 species and 1 species complex). In contrast, MALDI-TOF MS identified 74.78% (86/115) of the isolates, with an accuracy of 96.51% (83/86). The MS showed substantial agreement with ITS barcoding (κ = 0.65, p < 0.001) when all isolates were included, and this agreement strengthened (κ = 0.76, p < 0.001) when the analysis was restricted to species represented in the MS database. Furthermore, almost perfect agreement was observed (κ = 0.81, p < 0.001) for common pathogenic Candida species, achieving 100% identification rate and accuracy for C. albicans and C. tropicalis.
Conclusion: ITS barcoding remains the gold standard method for yeast identification. MALDI-TOF MS is a viable option for routine clinical use, but it requires database expansion and technical improvements for uncommon species or species complexes.
Keywords
DNA barcoding ITS MALDI-TOF MS Yeast identification
Fungal identification using conventional methods in routine practice often faces challenges, such as the need for culti-vation, being time-consuming, and low specificity. With the introduction of antifungal stewardship programs and routine antifungal susceptibility testing for managing fungal infections, precise identification of fungal pathogens has become in- creasingly important. Identification using DNA barcoding markers has been recognized as another "gold standard" method for fungal identification; however, this method entails high costs, complex protocols, and long turnaround times. The internal transcribed spacer (ITS) region of ribosomal DNA has been widely adopted as the primary DNA barcode for fungal identification due to several key attributes1,2, including the availability of comprehensive reference databases with type specimens to discriminate most species, standardized amplification protocols, and a phylogenetic framework that allows evolutionary interpretation. These features collectively establish ITS sequencing as the current reference standard for definitive fungal identification, particularly for species that cannot be reliably distinguished by phenotypic methods. Since MALDI-TOF MS was developed3,4, many studies have applied the MS to fungal identification, especially for yeast pathogens in routine diagnostics5-11, to overcome DNA barcoding dis- advantages. MALDI-TOF MS identifies microorganisms by analyzing their protein profiles, generating unique mass spectra patterns that serve as molecular fingerprints for each species. This technique has gained popularity in clinical microbiology due to its rapid turnaround time (minutes vs. days) and low operational costs12,13.
In the present study, we compared and assessed two molecular methods, MALDI-TOF MS and DNA barcoding using ITS, to identify yeast strains isolated from patients and domestic environments.
1. Collection of yeast strains
One hundred and fifteen yeast strains were collected from 2022 to 2025. Of these 89 strains (strains F1-F54 and B2-B57) were isolated from clinical specimens of patients who were admitted at the Srinagarind Hospital, Khon Kaen, Thailand. The remaining 26 strains (strains E3-E39) were collected from household environments within the Khon Kaen municipal area. The colonial morphology of each strain was examined using Sabouraud dextrose agar and CHROMagar (BrillianceTM Candida Agar, Oxoid, UK), while micromorphology was done using lactophenol cotton blue and/or nigrosin staining.
2. Identification by MALDI-TOF MS
All yeast strains were identified using MALDI-TOF MS (MALDI Biotyper®, autoflex® maX mass, Bruker Daltonics, Bremen, DE). The method was performed according to the manufacturer's protocol. Sample preparation followed the manufacturer's formic acid extraction protocol. Briefly, yeast cells were smeared onto a ground steel MALDI target plate (MTP 384), air-dried, overlaid with 1 μl of 70% formic acid and dried, then overlaid with 1 μl of α-cyano-4-hydroxycinnamic acid (HCCA) matrix solution and dried again. The steel plate was examined to obtain mass spectral profiles using an Autoflex maX mass spectrometer. Each yeast strain profile was identified using Bruker's MALDI Biotyper® Compass soft- ware version 4.1.100 with the MBT 8468 Species/Entry List Revision C (March 2019), and identifications were accepted according to the manufacturer's criteria.
3. PCR sequencing of ITS
All strains were DNA-extracted using a DNA extraction kit (Vivantis Technologies Sdn Bhd., Selangor Darul Ehsan, Malaysia). The DNAs were quantified using a NanoDrop 2000 spectrophotometer (Thermo Scientific, Massachusetts, USA). The ITS sequence in the ribosomal gene was amplified by PCR using the forward primers ITS1 (TCCGTAGGTGAACCTGCGG), V9G (TTACGTCCCTGCCCTTTGTA)14, or ITS5 (TCCTCCGCT- TATTGATATGC)15 and the reverse primers including ITS4 (GGAAGTAAAAGTCGTAACAAGG)15 or LS266 (GCATTCCC- AAACAACTCGACTC)16. Different primer combinations were used to optimize amplification success across diverse yeast taxa. The primer pair ITS1/ITS4 was primarily used, while alter- native primer pairs (V9G/LS266 or ITS5/ITS4) were employed when amplification failed or produced weak products. The PCR reaction contained 10 μM of each primer, 12.5 μl of KOD One® PCR Master Mix (TOYOBO, JP), and >10 ng genomic fungal DNA in 25 μl of total volume reaction. The PCR conditions started at 95℃ for 5 min, followed by 35 cycles of denaturation at 95℃ for 35 sec, annealing at 48℃ for 30 sec, and extension at 72℃ for 30 sec. Amplification was performed using a Veriti® Thermal Cycler (Applied Biosystems, CA, USA). The amplification products were examined with 2% gel electrophoresis and submitted to Sanger's sequencing.
4. Identification by ITS barcoding
The ITS sequences obtained from Sanger's sequencing were edited using BIOEDIT v.7.2.517 and blasted against the type materials in GenBank using megablast in BLAST® blastn18. The datasets, including the sequences of the tested strains and those retrieved from megablast, were prepared using BIOEDIT v.7.2.5 and aligned with the algorithms, specifically ClustalW in BIOEDIT v.7.2.5, Multiple Alignment using Fast Fourier Transform (MAFFT)19, or MUltiple Sequence Com- parison by Log-Expectation (MUSCLE)20 via the EMBI-EBI web server21. Phylogenetic analyses for species identification were performed using MEGA1122. The best model and appro- priate phylogenetic algorithms were inferred to reconstruct a phylogenetic tree and to estimate percentage similarity. Species identification was considered valid when the test strain formed a monophyletic clade with the type species in the phylogenetic tree and/or showed sequence similarity with the type specimen across the ITS region.
5. Assessment of MALDI-TOF MS for yeast identification
With ITS barcoding serving as the reference method, the identification rate and accuracy of MALDI-TOF MS were cal- culated using the following formulas:
Agreement between the two methods was assessed using Cohen's kappa coefficient, calculated for two scenarios: (1) all isolates and (2) isolates of species represented in the MALDI-TOF MS database. Statistical analysis was performed using SPSS version 29.0 (IBM Corp., Armonk, NY, USA). The Cohen's kappa (κ) coefficients were interpreted based on the criteria established by Landis and Koch23: ≤ 0 (no agreement), 0.01-0.20 (slight), 0.21-0.40 (fair), 0.41-0.60 (moderate), 0.61-0.80 (substantial), and 0.81-1.00 (almost perfect agree- ment).
1. Identification by MALDI-TOF MS
MALDI-TOF MS identified 86 strains belonging to 11 species (number of strains), including C. albicans (46), C. tropicalis (22), C. metapsilosis (2), C. parapsilosis (1), Meyerozyma guil- liermondii (syn. Candida guilliermondii) (1), Nakaseomyces glabratus (syn. Candida glabrata) (7), Pichia kudriavzevii (syn. Candida krusei) (2), Diutina rugosa (syn. Candida rugosa) (1), Kodamaea ohmeri (1), Cystobasidium minutum (syn. Rhodotorula minuta) (1), and Rhodotorula mucilaginosa (1). However, the MS failed to identify 29 strains. All results are shown in Table 1.
2. Identification by ITS barcoding
All 115 yeast strains were identified using phylogenetic analysis of their ITS sequences in comparison with the type materials. Species identification was determined based on monophyletic clustering and/or the highest sequence similarity with the type species, as shown in Fig. 1. Of the 115 strains, 102 were classified as ascomycetous yeasts belonging to 14 species, whereas 13 strains were classified as basidiomycetous yeasts, comprising 5 species and 1 species complex. The ascomycetous species (number of strains) included Candida albicans (46), C. tropicalis (22), Nakaseomyces glabratus (8), C. orthopsilosis (6), C. metapsilosis (4), Pichia kudriavzevii (4), Wickerhamomyces sydowiorum (3), C. parapsilosis (2), Debaryomyces nepalensis (2), Meyerozyma caribbica (1), Kodamaea ohmeri (1), Diutina mesorugosa (1), Trichomona- scus ciferrii (syn. Candida ciferrii) (1), and Yarrowia lipolytica (1). The basidiomycetous sp ecies (number of strains) included Papiliotrema rajasthanensis (5), Rhodotorula mucilaginosa (3), Kwoniella heveanensis (2), Cystobasidium benthicum (1), Cystobasidium minutum (1), and Naganishia albidosimilis-liquefaciens (syn. Cryptococcus albidosimilis and C. lique- faciens) (1). Candida albicans, C. tropicalis, C. parapsilosis complex, and Nakaseomyces glabratus were the predominant species isolated from patients' specimens, whereas basidio- mycetous species predominated in environmental specimens. All details are presented in Table 1.
|
No. |
Strain |
Source |
Maldi-TOF MS |
ITS barcoding |
|
1 |
E3 |
Refrigerator |
No identification possible |
Candida
orthopsilosis |
|
2 |
E5 |
Electric fan |
Cystobasidium
minutum |
Cystobasidium
minutum |
|
3 |
E6 |
Refrigerator |
No identification possible |
Papiliotrema
rajasthanensis |
|
4 |
E8 |
Air conditioner |
No identification possible |
Naganishia
albidosimilis-liquefaciens |
|
5 |
E9 |
Air conditioner |
No identification possible |
Papiliotrema
rajasthanensis |
|
6 |
E11 |
Floor |
No identification possible |
Candida
metapsilosis |
|
7 |
E12 |
Floor |
No identification possible |
Rhodotorula
mucilaginosa |
|
8 |
E13 |
Lid of trash box |
No identification possible |
Candida
orthopsilosis |
|
9 |
E14 |
Air purifier |
Candida
metapsilosis |
Candida metapsilosis |
|
10 |
E15 |
Refrigerator |
No identification possible |
Rhodotorula
mucilaginosa |
|
11 |
E17 |
Bathroom |
No identification possible |
Wickerhamomyces
sydowiorum |
|
12 |
E19 |
Air conditioner |
No identification possible |
Debaryomyces
nepalensis |
|
13 |
E21 |
Office area |
No identification possible |
Candida
orthopsilosis |
|
14 |
E22 |
Bathroom |
No identification possible |
Wickerhamomyces
sydowiorum |
|
15 |
E23 |
Electric fan |
No identification possible |
Candida
metapsilosis |
|
16 |
E25 |
Glass window |
No identification possible |
Papiliotrema
rajasthanensis |
|
17 |
E26 |
Bathroom |
No identification possible |
Wickerhamomyces
sydowiorum |
|
18 |
E27 |
Air conditioner |
No identification possible |
Cystobasidium
benthicum |
|
19 |
E28 |
Floor |
No identification possible |
Yarrowia
lipolytica |
|
20 |
E29 |
Door |
No identification possible |
Kwoniella
heveanensis |
|
21 |
E31 |
Table |
No identification possible |
Papiliotrema
rajasthanensis |
|
22 |
E32 |
Bathroom |
Rhodotorula
mucilaginosa |
Rhodotorula
mucilaginosa |
|
23 |
E33 |
Air conditioner |
Candida
parapsilosis |
Candida
parapsilosis |
|
24 |
E34 |
Floor |
No identification possible |
Candida
orthopsilosis |
|
25 |
E36 |
Glass window |
No identification possible |
Papiliotrema
rajasthanensis |
|
26 |
E39 |
Door |
No identification possible |
Kwoniella
heveanensis |
|
27 |
F1 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
28 |
F2 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
29 |
F3 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
30 |
F4 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
31 |
F5 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
32 |
F6 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
33 |
F7 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
34 |
F8 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
35 |
F9 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
36 |
F12 |
Patient's fluid |
Candida
albicans |
Candida albicans |
|
37 |
F13 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
38 |
F15 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
39 |
F16 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
40 |
F18 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
41 |
F19 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
42 |
F20 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
43 |
F22 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
44 |
F25 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
45 |
F26 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
46 |
F27 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
47 |
F28 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
48 |
F29 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
49 |
F30 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
50 |
F31 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
51 |
F32 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
52 |
F33 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
53 |
F34 |
Patient's fluid |
Candida
albicans |
Candida
albicans |
|
54 |
F35 |
Patient's fluid |
No identification possible |
Pichia
kudriavzevii |
|
55 |
F36 |
Patient's fluid |
No identification possible |
Candida
orthopsilosis |
|
56 |
F38 |
Patient's fluid |
No identification possible |
Nakaseomyces
glabratus |
|
57 |
F40 |
Patient's fluid |
No identification possible |
Trichomonascus
ciferrii |
|
58 |
F41 |
Patient's fluid |
No identification possible |
Candida
parapsilosis |
|
59 |
F43 |
Patient's fluid |
Pichia
kudriavzevii |
Pichia
kudriavzevii |
|
60 |
F47 |
Patient's fluid |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
61 |
F48 |
Patient's fluid |
Diutina
rugosa |
Diutina
mesorugosa |
|
62 |
F49 |
Patient's fluid |
Candida
metapsilosis |
Candida
metapsilosis |
|
63 |
F50 |
Patient's fluid |
Candida
tropicalis |
Candida
tropicalis |
|
64 |
F51 |
Patient's fluid |
Candida tropicalis |
Candida
tropicalis |
|
65 |
F52 |
Patient's fluid |
Candida
tropicalis |
Candida
tropicalis |
|
66 |
F53 |
Patient's fluid |
Candida
tropicalis |
Candida
tropicalis |
|
67 |
F54 |
Thrombus |
Candida
tropicalis |
Candida
tropicalis |
|
68 |
B2 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
69 |
B4 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
70 |
B5 |
Hemoculture |
Kodamaea
ohmeri |
Kodamaea
ohmeri |
|
71 |
B6 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
72 |
B7 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
73 |
B8 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
74 |
B9 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
75 |
B10 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
76 |
B12 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
77 |
B13 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
78 |
B14 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
79 |
B15 |
Hemoculture |
Candida tropicalis |
Candida
tropicalis |
|
80 |
B16 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
81 |
B19 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
82 |
B20 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
83 |
B21 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
84 |
B22 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
85 |
B23 |
Hemoculture |
Candida
tropicalis |
Candida
tropicalis |
|
86 |
B26 |
Hemoculture |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
87 |
B27 |
Hemoculture |
No identification possible |
Pichia
kudriavzevii |
|
88 |
B28 |
Hemoculture |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
89 |
B29 |
Hemoculture |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
90 |
B31 |
Hemoculture |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
91 |
B33 |
Hemoculture |
Candida
parapsilosis |
Candida
orthopsilosis |
|
92 |
B35 |
Hemoculture |
Meyerozyma
guilliermondii |
Meyerozyma
caribbica |
|
93 |
B36 |
Hemoculture |
No identification possible |
Debaryomyces
nepalensis |
|
94 |
B37 |
Hemoculture |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
95 |
B38 |
Hemoculture |
Pichia
kudriazevii |
Pichia
kudriazevii |
|
96 |
B39 |
Hemoculture |
Nakaseomyces
glabratus |
Nakaseomyces
glabratus |
|
97 |
B43 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
98 |
B44 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
99 |
B45 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
100 |
B46 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
101 |
B47 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
102 |
B48 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
103 |
B49 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
104 |
B50 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
105 |
B51 |
Hemoculture |
Candida
albicans |
Candida
albicans |
|
106 |
B52 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
107 |
B53 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
108 |
B54 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
109 |
B55 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
110 |
B56 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
111 |
B57 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
112 |
B58 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
113 |
B59 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
114 |
B60 |
Hemoculture |
Candida
albicans |
Candida albicans |
|
115 |
B61 |
Hemoculture |
Candida
albicans |
Candida albicans |
3. Assessment of MALDI-TOF MS for yeast identification
MALDI-TOF MS and ITS barcoding results are compared in Table 1. In addition to the 29 unidentified strains, the MS misidentified three strains as Candida parapsilosis, Diutina rugosa, and Meyerozyma guilliermondii; these were sub- sequently identified as C. orthopsilosis, D. mesorugosa, and M. caribbica, respectively, based on ITS sequencing. Using ITS barcoding as the reference method, the overall identification rate for MALDI-TOF MS was 74.78%, with an overall accuracy of 96.51%. The MS system demonstrated the highest identi- fication rate for Candida species (100%), whereas the lowest rate was observed for basidiomycetous species (15.38%). Specifically, accuracy values were 98.61% for Candida species, 83.33% for non-Candida species, and 100% for basidio- mycetous species. Cohen's kappa analysis demonstrated substantial agreement between ITS barcoding and MALDI-TOF MS across all isolates (n = 115, κ = 0.65, p < 0.001). This agreement strengthened when the analysis was restricted to isolates of species represented in the MALDI-TOF MS database (n = 100, κ = 0.76, p < 0.001). Moreover, almost perfect agreement (κ = 0.81, p < 0.001) was achieved for common pathogenic Candida species, including C. albicans, C. tropicalis, and the C. parapsilosis complex. Detailed results are shown in Table 2.
| Cluster/ Group |
Species | Source‡ (B/F/E) |
ITS | MALDI-TOF MS | Cohen's kappa |
|||||
|---|---|---|---|---|---|---|---|---|---|---|
| Right Ident. | Mis. Ident. | Not Ident. | Ident. (%) | Accur. (%) | ||||||
| Ascomycetous yeasts | Candida species | Candida albicans | 19/27/- | 46 | 46 | 0 | 0 | 100 | 100 | |
| C. tropicalis | 17/5/- | 22 | 22 | 0 | 0 | 100 | 100 | |||
| C. parapsilosis | -/1/1 | 2 | 1 | 0 | 1 | 50 | 100 | |||
| C. metapsilosis | -/1/3 | 4 | 2 | 0 | 2 | 50 | 100 | |||
| C. orthopsilosis | 1/1/4 | 6 | 0 | 1 | 5 | 16.66 | 0 | |||
| Total number of each source | 37/35/8 | |||||||||
| MALDI-TOF assessment | ||||||||||
| Only species in MS-database | 80 | 71 | 1 | 8 | 90.00 | 98.61 | 0.81§ | |||
| All species | 80 | 71 | 1 | 8 | 90.00 | 98.61 | 0.81§ | |||
| Non-Candida species | Meyerozyma caribbica* | 1/-/- | 1 | 0 | 1 | 0 | 100 | 0 | ||
| Debaryomyces nepalensis* | 1/-/1 | 2 | 0 | 0 | 2 | 0 | - | |||
| Wickerhamomyces sydowiorum* | -/-/3 | 3 | 0 | 0 | 3 | 0 | - | |||
| Kodamaea ohmeri | 1/-/- | 1 | 1 | 0 | 0 | 100 | 100 | |||
| Diutina mesorugosa* | -/1/- | 1 | 0 | 1 | 0 | 100 | 0 | |||
| Pichia kudriavzevii | 2/2/- | 4 | 2 | 0 | 2 | 50 | 100 | |||
| Trichomonascus ciferrii | -/1/- | 1 | 0 | 0 | 1 | 0 | - | |||
| Nakaseomyces glabatus | 6/2/- | 8 | 7 | 0 | 1 | 87.50 | 100 | |||
| Yarrowia lipolytica | -/-/1 | 1 | 0 | 0 | 1 | 0 | - | |||
| Total number of each source | 11/6/5 | |||||||||
| MALDI-TOF assessment | ||||||||||
| Only species in MS-database | 15 | 10 | 0 | 5 | 66.66 | 100 | 0.53§ | |||
| All species | 22 | 10 | 2 | 10 | 54.54 | 83.33 | 0.37§ | |||
| Basidiomycetous yeasts | Cystobasidium benthicum* | -/-/1 | 1 | 0 | 0 | 1 | 0 | - | ||
| Cystobasidium minutum | -/-/1 | 1 | 1 | 0 | 0 | 100 | 100 | |||
| Rhodotorula mucilaginosa | -/-/3 | 3 | 1 | 0 | 2 | 33.33 | 100 | |||
| Naganishia albidosimilis-liquefaciens† | -/-/1 | 1 | 0 | 0 | 1 | 0 | - | |||
| Kwoniella heveanensis* | -/-/2 | 2 | 0 | 0 | 2 | 0 | - | |||
| Papiliotrema rajasthanensis* | -/-/5 | 5 | 0 | 0 | 5 | 0 | - | |||
| Total number of each source | -/-/13 | |||||||||
| MALDI-TOF assessment | ||||||||||
| Only species in MS-database | 5 | 2 | 0 | 3 | 40 | 100 | 0.28∥ | |||
| All species | 13 | 2 | 0 | 11 | 15.38 | 100 | 0.13§ | |||
| Grand Total | Total number of each source | 48/41/26 | ||||||||
| Total MALDI-TOF assessment | ||||||||||
| Only species in MS-database | 100 | 83 | 1 | 16 | 84 | 98.81 | 0.76§ | |||
| All species | 115 | 83 | 3 | 29 | 74.78 | 96.51 | 0.65§ | |||
|
* = Not in MS-database, † = species complex, ‡ = Source of strain: B (Blood)/F (Fluid&other)/E (Domestic environment) § = p < 0.001, ∥ = p < 0.05 |
||||||||||
Invasive fungal infections, particularly candidemia and in- vasive aspergillosis, are associated with mortality rates of 20-40% and 30-95%, respectively, depending on the patient population and timeliness of appropriate therapy24,25. Early identification has been shown to improve survival outcomes26. Recent stewardship studies have demonstrated that rapid diagnosis can reduce inappropriate antifungal use and shorten hospital stays27.
In our study, DNA barcoding using ITS yielded satisfactory results, confirming its role as another "gold standard" method for yeast identification. The well-supported phylogenetic species clusters and/or the minimal genetic distances to type strains serve as the criteria for sequence-based identification, as previously mentioned28,29. DNA barcoding represents a highly robust methodology for species identification, as its procedures align with the foundational protocols established in contemporary yeast taxonomy through molecular phylo- genetic analysis. However, to ensure precise identification, DNA barcoding necessitates the integration of molecular techniques and deep taxonomic knowledge. While BLAST® blastn18 appears to be an efficient tool for identifying DNA barcoding sequences, results can sometimes be difficult to interpret without sufficient expertise. Although modern fungal taxonomy relies on the genealogical species concept, challenges persist regarding taxonomic non-consensus and species complexes. Assigning a single barcoding marker often proves problematic for species defined via multigene analysis or those with ambiguous classifications. Therefore, phylo- genetic analysis, including comparisons with type materials, is essential to achieve accurate identification. Our study utilized a straightforward phylogenetic approach using ITS, a method rarely incorporated into MALDI-TOF assessment studies. This approach successfully identified nearly all patho- genic species at both the species and species complex levels. Notably, ITS cannot differentiate two basidiomycetous species, Naganishia albidosimilis and N. liquefaciens (syn. Cryptococcus albidosimilis and C. liquefaciens), due to their identical ITS sequences. However, these species can be distinguished through phylogenetic analysis of an alternative marker, the D1/D2 domain of the LSU rDNA30. Despite its high resolution, DNA barcoding remains limited by long turnaround times and high operational costs, which are significant drawbacks for routine clinical use.
Across all species investigated, MALDI-TOF MS showed sub- stantial agreement with ITS barcoding (κ = 0.65), with an identification rate of 74.78% and a high accuracy of 96.51%. However, the system failed to identify seven species not represented in its database, highlighting a primary limitation of MALDI-TOF MS. These species included Meyerozyma caribbica, Debaryomyces nepalensis, Wickerhamomyces sydo- wiorum, Diutina mesorugosa, Cystobasidium benthicum, Kwoniella heveanensis, and Papiliotrema rajasthanensis. When the analysis was restricted to species represented in the database, the MS demonstrated higher agreement with ITS barcoding (κ = 0.76), with the identification rate and accuracy reaching 84% and 98.91%, respectively. Identification rates for uncommon yeasts and molds typically range from 65% to 85%, depending on the instrument and database10. Although the MS database used in this study included many species (Table 2), it still failed to identify certain strains. This may be attributed to factors such as the partial extraction method used for the specimens31 or strain variations in mass spectral profiles32.
The overall identification accuracy in our study was highly satisfactory at 96.51%, reaching 98.81% when restricted to species present in the database. Such high accuracy (often > 95%) is consistent with findings for common yeast identification using either the Bruker MALDI Biotyper or the bioMérieux VITEK MS/MS33-35. MALDI-TOF MS, when com- bined with curated libraries or purpose-built datasets, has been shown to successfully differentiate species complexes, including C. parapsilosis, C. guilliermondii, C. glabrata, and Cryptococcus neoformans/gattii 8-11.
However, in this study, the MS failed to differentiate the species within the C. parapsilosis complex and the Meyero- zyma guilliermondii (syn. Candida guilliermondii) complex. In addition to technical limitations, incomplete databases, and variations in mass spectral profiles, the close genetic relation- ships among these species could be a plausible explanation for identification failure. MALDI-TOF MS identifies organisms based on ribosomal protein mass spectra, which are highly conserved among closely related species36. When species share more than 98% sequence similarity, discriminatory protein peaks may be subtle or absent, resulting in misidentification or non-identification. This phenomenon has been documented in other fungal complexes, including Aspergillus niger, A. flavus, and the A. nidulans complex37, as well as the Crypto- coccus gattii complex11. Similarly, the close phylogenetic relationship between C. albicans and C. africana poses signifi- cant challenges for MALDI-TOF MS38. Database expansion alone may not fully resolve this issue. Species complexes with minimal protein-level divergence may require supplementary identification methods, such as ITS sequencing or multilocus sequence typing, for definitive differentiation39.
From a clinical perspective, these misidentifications may have limited therapeutic impact. Species within the C. para- psilosis complex generally exhibit similar antifungal suscepti- bility profiles, with minimal differences in minimum inhibitory concentration distributions for azoles and echinocandins40,41. Similarly, M. caribbica and M. guilliermondii show comparable susceptibility patterns42. No inter-complex misidentifications that could lead to inappropriate antifungal selection were observed in our cohort. However, accurate species-level iden- tification within complexes remains important for epidemio- logical tracking and understanding virulence differences. For instance, Candida orthopsilosis and C. metapsilosis differ from C. parapsilosis in biofilm formation capacity and tissue tropism43. Such information is valuable for infection control and outbreak investigation, even when immediate therapeutic decisions are unaffected.
In our findings, when focusing on the most common patho- genic Candida species present in the database, MALDI-TOF MS showed almost perfect agreement with ITS barcoding (κ = 0.81), achieving 100% identification rate and accuracy in C. albicans and C. tropicalis. Furthermore, the MS proved highly reliable in identifying Nakaseomyces glabratus (syn. Candida glabrata) with 100% accuracy. To improve the identification capability and accuracy of MALDI-TOF MS, expanding the database to include a broader range of species associated with human and animal infections, such as Meyerozyma caribbica42,44, Diutina mesorugosa45, and Debaryomyces nepalensis46, should be considered a critical priority.
Additionally, our findings revealed that members of the C. parapsilosis complex, viz., C. parapsilosis, C. orthopsilosis, and C. metapsilosis, were isolated from various environmental surfaces, including floors, electric fans, air conditioners, air purifiers, trash bins, and refrigerators. These observations are consistent with their established environmental persistence and hand-mediated transmission pathways47. These results highlight the critical role of environmental surveillance in monitoring the dissemination of azole-resistant strains48 or potential sources of infection49.
In conclusion, both MALDI-TOF MS and ITS barcoding have distinct roles in clinical mycology laboratories. For cost-effective laboratory practice, we recommend MALDI-TOF MS as the first-line method for rapid identification of com- mon yeast pathogens. ITS barcoding should be reserved for unidentified isolates, species complexes, rare species, and high-impact clinical cases requiring definitive identification39,50-52. Continued expansion of MALDI-TOF reference databases, particularly to include closely related or cryptic species as well as newly described species or discovered pathogens, will further enhance its utility.
References
1. Perlin DS, Wiederhold NP. Culture-independent molecular methods for detection of antifungal resistance mech- anisms and fungal identification. J Infect Dis 2017;216: S458-S465
Google Scholar
2. Wickes BL, Wiederhold NP. Molecular diagnostics in medical mycology. Nat Commun 2018;9:5135
Google Scholar
3. Karas M, Bachmann D, Hillenkamp F. Influence of the wavelength in high-irradiance ultraviolet laser desorption mass spectrometry of organic molecules. Anal Chem 1985;57:2935-2939
Google Scholar
4. Tanaka K, Waki H, Ido Y, Akita S, Yoshida Y, Yoshida T, et al. Protein and polymer analyses up to m/z 100 000 by laser ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 1988;2:151-153
Google Scholar
5. Patel R. A moldy application of MALDI: MALDI-ToF mass spectrometry for fungal identification. J Fungi 2019;5:4
Google Scholar
6. Angeletti S, Ciccozzi M. Matrix-assisted laser desorption ionization time-of-flight mass spectrometry in clinical microbiology: An updating review. Infect Genet Evol 2019;76:104063
Google Scholar
7. Walsh TJ, McCarthy MW. The expanding use of matrix-assisted laser desorption/ionization-time of flight mass spectroscopy in the diagnosis of patients with mycotic diseases. Expert Rev Mol Diagn 2019;19:241-248
Google Scholar
8. Hou X, Xiao M, Chen SCA, Wang H, Yu SY, Fan X, et al. Identification and antifungal susceptibility profiles of Candida nivariensis and Candida bracarensis in a multi-center Chinese collection of yeasts. Front Microbiol 2017: 8
Google Scholar
9. Samantaray S, Singh R. Evaluation of MALDI-TOF MS for identification of species in the Candida parapsilosis complex from candidiasis cases. J Appl Lab Med 2022; 7:889-900
Google Scholar
10. Dutkiewicz M, Garros M, Bui J, Charlier V, Da Silva E, Lemaire M, et al. Comparison of MALDI-TOF MS instru- ments and databases for the identification of uncommon yeasts, Aspergillus spp. and rare filamentous fungi. J Clin Microbiol 2025;63:e01612-624
Google Scholar
11. Bernhard M, Worasilchai N, Kangogo M, Bii C, Trzaska WJ, Weig M, et al. CryptoType - Public datasets for MALDI-TOF-MS based differentiation of Cryptococcus neoformans/gattii complexes. Front Cell Infect Microbiol 2021;11:634382
Google Scholar
12. Tan KE, Ellis BC, Lee R, Stamper PD, Zhang SX, Carroll KC. Prospective evaluation of a matrix-assisted laser desorption ionization-time of flight mass spectrometry system in a hospital clinical microbiology laboratory for identification of bacteria and yeasts: A bench-by-bench study for assessing the impact on time to identification and cost-effectiveness. J Clin Microbiol 2012;50:3301-3308
Google Scholar
13. Robert MG, Cornet M, Hennebique A, Rasamoelina T, Caspar Y, Pondérand L, et al. MALDI-TOF MS in a medical mycology laboratory: On stage and backstage. Microorganisms 2021;9:1283
Google Scholar
14. de Hoog GS, Gerrits van den Ende AH. Molecular diag- nostics of clinical strains of filamentous Basidiomycetes. Mycoses 1998;41:183-189
Google Scholar
15. White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR Protocols: A guide to methods and applications. New York: Academic Press, 1989:315-322
Google Scholar
16. Masclaux F, Guého E, de Hoog GS, Christen R. Phylo- genetic relationships of human-pathogenic Cladosporium (Xylohypha) species inferred from partial LS rRNA sequences. J Med Vet Mycol 1995;33:327-338
Google Scholar
17. Hall TA. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In: Nucleic acids symposium series no. 41. Oxford University Press, 1999:95-98
Google Scholar
18. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990;215: 403-410
Google Scholar
19. Katoh K, Standley DM. MAFFT multiple sequence align- ment software version 7: Improvements in performance and usability. Mol Biol Evol 2013;30:772-780
Google Scholar
20. Edgar RC. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 2004;32:1792-1797
Google Scholar
21. Madeira F, Madhusoodanan N, Lee J, Eusebi A, Niewielska A, Tivey ARN, et al. The EMBL-EBI job Dis- patcher sequence analysis tools framework in 2024. Nucleic Acids Res 2024;52:W521-W525
Google Scholar
22. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-3027
Google Scholar
23. Landis JR, Koch GG. The measurement of observer agreement for categorical data. Biometrics 1977;33: 159-174
Google Scholar
24. World Health Organization. WHO fungal priority patho- gens list to guide research, development and public health action. 1st ed. Geneva: World Health Organization; 2022
Google Scholar
25. Fisher MC, Alastruey-Izquierdo A, Berman J, Bicanic T, Bignell EM, Bowyer P, et al. Tackling the emerging threat of antifungal resistance to human health. Nat Rev Microbiol 2022;20:557-571
Google Scholar
26. Paramythiotou E, Frantzeskaki F, Flevari A, Armaganidis A, Dimopoulos G. Invasive fungal infections in the ICU: How to approach, how to treat. Molecules 2014;19: 1085-1119
Google Scholar
27. Mutepfa CC, Suklan J, Bell J, Guiney M, Jones W, Simpson J, et al. Rapid diagnostic tests to inform clinical decision-making for antifungal stewardship in the ICU: A qualitative study with NHS staff, patients, and their legal representatives. Health Technol Assess 2025:1-19
Google Scholar
28. Meyer CP, Paulay G. DNA barcoding: Error rates based on comprehensive sampling. PLoS Biol 2005;3:e422
Google Scholar
29. Collins RA, Cruickshank RH. The seven deadly sins of DNA barcoding. Mol Ecol Resour 2013;13:969-975
Google Scholar
30. Conde-Pereira C, Rodas-Rodríguez L, Díaz-Paz M, Palacios-Rivera H, Firacative C, Meyer W, et al. Fatal case of polymicrobial meningitis caused by Cryptococcus liquefaciens and Mycobacterium tuberculosis complex in a human immunodeficiency virus-infected patient. J Clin Microbiol 2015;53:2753-2755
Google Scholar
31. Fraser M, Brown Z, Houldsworth M, Borman AM, Johnson EM. Rapid identification of 6328 isolates of pathogenic yeasts using MALDI-ToF MS and a simplified, rapid extraction procedure that is compatible with the Bruker Biotyper platform and database. Med Mycol 2016;54: 80-88
Google Scholar
32. Moothoo-Padayachie A, Kandappa HR, Krishna SBN, Maier T, Govender P. Biotyping Saccharomyces cerevisiae strains using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS). Eur Food Res Technol 2013;236:351-364
Google Scholar
33. Teke L, Barış A, Bayraktar B. Comparative evaluation of the Bruker Biotyper and Vitek MS matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) systems for non-albicans Candida and uncommon yeast isolates. J Microbiol Methods 2021; 185:106232
Google Scholar
34. Dichtl K, Klugherz I, Greimel H, Luxner J, Köberl J, Friedl S, et al. A head-to-head comparison of three MALDI-TOF mass spectrometry systems with 16S rRNA gene sequencing. J Clin Microbiol 2023;61:e0191322
Google Scholar
35. Hong K, Bae J, Lee H, Yong D, Cho HW, Kim YA. A head-to-head comparison of five MALDI-TOF mass spectro- metry systems for detection of clinically relevant bacteria and fungi, a single center study. J Glob Antimicrob Resist 2024;39:37
Google Scholar
36. Singhal N, Kumar M, Kanaujia PK, Virdi JS. MALDI-TOF mass spectrometry: An emerging technology for microbial identification and diagnosis. Front Microbiol 2015;6:791
Google Scholar
37. Shao J, Wang Q, Wei L, Wan Z, Li R, Yu J. Limitations of matrix-assisted laser desorption/ionization time-of-flight mass spectrometry for the identification of Aspergillus species. Med Mycol 2022;60:myab084
Google Scholar
38. Hazirolan G, Altun HU, Gumral R, Gursoy NC, Otlu B, Sancak B. Prevalence of Candida africana and Candida dubliniensis, in vulvovaginal candidiasis: First Turkish Candida africana isolates from vulvovaginal candidiasis. J Mycol Med 2017;27:376-381
Google Scholar
39. Morovati H, Kord M, Ahmadikia K, Eslami S, Hemmatzadeh M, Kurdestani KM, et al. A comprehensive review of identification methods for pathogenic yeasts: Challenges and approaches. Adv Biomed Res 2023;12: 187
Google Scholar
40. Guo J, Zhang M, Qiao D, Shen H, Wang L, Wang D, et al. Prevalence and antifungal susceptibility of Candida parapsilosis species complex in Eastern China: A 15-year retrospective study by ECIFIG. Front Microbiol 2021;12: 644000
Google Scholar
41. Zhang W, Zhan M, Wang N, Fan J, Han X, Li C, et al. In vitro susceptibility profiles of Candida parapsilosis species complex subtypes from deep infections to nine antifungal drugs. J Med Microbiol 2023;72:001640
Google Scholar
42. Ghasemi R, Lotfali E, Rezaei K, Madinehzad SA, Tafti MF, Aliabadi N, et al. Meyerozyma guilliermondii species complex: Review of current epidemiology, antifungal resistance, and mechanisms. Braz J Microbiol 2022;53: 1761-1779
Google Scholar
43. Lockhart SR, Messer SA, Pfaller MA, Diekema DJ. Geographic distribution and antifungal susceptibility of the newly described species Candida orthopsilosis and Candida metapsilosis in comparison to the closely related species Candida parapsilosis. J Clin Microbiol 2008;46: 2659-2664
Google Scholar
44. Chaves ALS, Trilles L, Alves GM, Figueiredo-Carvalho MHG, Brito-Santos F, Coelho RA, et al. A case-series of bloodstream infections caused by the Meyerozyma guilliermondii species complex at a reference center of oncology. Brazil Med Mycol 2021;59:235-243
Google Scholar
45. Adjapong G, Bartlett M, Hale M, Garrill A. The isolation of Candida rugosa and Candida mesorugosa from clinical samples in Ghana. Med Mycol 2016;54:322-326
Google Scholar
46. Moretti A, Fukushima K, Takizawa K, Suzuki M, Vidotto V, Cannizzo FT, et al. First report of oral colonization by Debaryomyces nepalensis in a dog. Mycopathologia 2007;164:189-192
Google Scholar
47. Trofa D, Gácser A, Nosanchuk JD. Candida parapsilosis, an emerging fungal pathogen. Clin Microbiol Rev 2008; 21:606-625
Google Scholar
48. Thomaz DY, de Almeida Jr JN, Sejas ONE, Del Negro GMB, Carvalho GOMH, Gimenes VMF, et al. Environ- mental clonal spread of azole-resistant Candida para- psilosis with Erg11-Y132F mutation causing a large candidemia outbreak in a Brazilian Cancer Referral Center. J Fungi (Basel) 2021;7:259
Google Scholar
49. Guinea J, Escribano P, Cadeau M, Lombardi L, Morio F. Emerging antifungal resistance in Candida parapsilosis: The end of the innocence. npj Antimicrob Resist 2025; 3:99
Google Scholar
50. Aydin M, Kustimur S, Kalkanci A, Duran T. Identification of medically important yeasts by sequence analysis of the internal transcribed spacer and D1/D2 region of the large ribosomal subunit. Rev Iberoam Micol 2019;36: 129-138
Google Scholar
51. Nascimento T, Inácio J, Guerreiro D, Diaz P, Patrício P, Proença L, et al. Enhancing ICU Candida spp. surveillance: A cost-effective approach focused on Candida auris detection. Front Cell Infect Microbiol 2024;14:1463456
Google Scholar
52. Hong H, Ximing Y, Jinghan M, Al-danakh A, Shujuan P, Ying L, et al. Candida auris infection; diagnosis, and resistance mechanism using high-throughput sequencing technology: A case report and literature review. Front Cell Infect Microbiol 2023;13:1211626
Google Scholar
Congratulatory MessageClick here!